tri3_tri6
Below is a demonstration of the features of the tri3_tri6 function
Contents
Syntax
[TRI6,V6,VX6C]=tri3_tri6(TRI3,V3,VXC);
Description
The tri3_tri6 converts 3-node triangles to 6-node triangles. These correspond to linear and quadratic triangular elements for finite element analysis. The 6-node triangular element follows the FEBio format such that the first 3 points are the 3-node triangle points, the following 3 nodes are the mid-edge points. To plot it with a command such as patch or gpatch one therefore needs to use element2patch or use: TRI6(:,[1 4 2 5 3 6]);
Examples
clear; close all; clc; % Plot settings fontSize=25; faceColor='b'; faceAlpha=0.3; edgeColor='k'; edgeWidth1=2; edgeWidth2=1; markerSize1=75; markerSize2=10;
CONVERSION FROM TRI3 TO TRI6
Creating an example triangulated mesh
[TRI3,V3]=geoSphere(2,1);
Converting to a single 10-node tetrahedron
[TRI6,V6,~]=tri3_tri6(TRI3,V3);
[F6]=element2patch(TRI6,[],'tri6');
Plotting elements
cFigure; % Open figure for plotting subplot(1,2,1); hold on; title('3-node linear triangles','FontSize',fontSize); gpatch(TRI3,V3,'rw'); %Plotting surface patchNormPlot(TRI3,V3); %Plotting face normals plotV(V3,'k.','MarkerSize',25); axisGeom; camlight('headlight'); subplot(1,2,2); hold on; title('6-node quadratic triangles','FontSize',fontSize); % gpatch(TRI6(:,[1 4 2 5 3 6]),V6,'gw'); %Plotting surface gpatch(F6,V6,'gw'); %Plotting surface patchNormPlot(F6,V6); %Plotting face normals plotV(V6,'k.','MarkerSize',25); axisGeom; camlight('headlight'); drawnow;
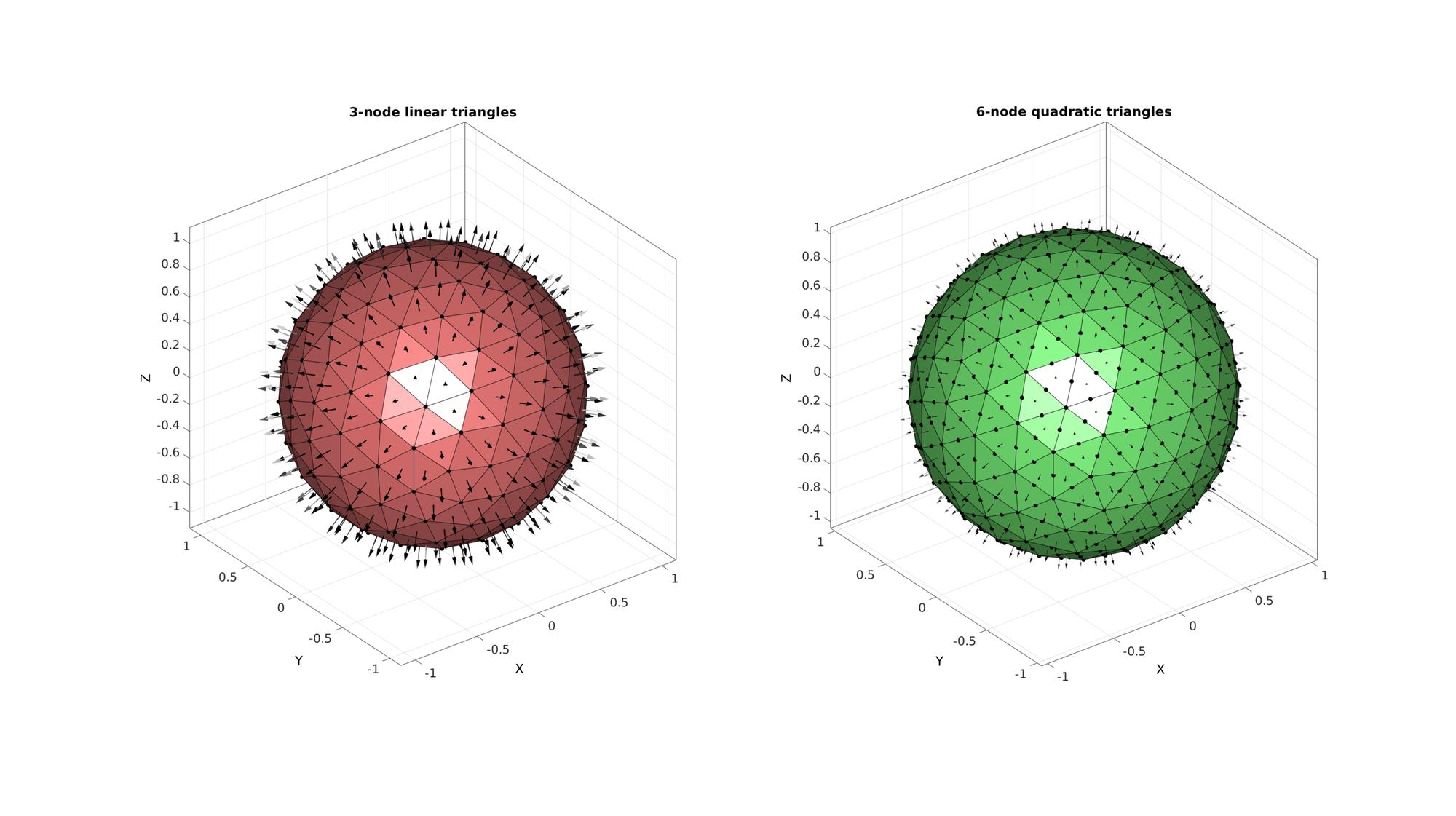
CONVERSION FROM TRI3 TO TRI6, EXAMPLE FOR A CONVERSION OF NODAL PARAMETERS
nodalData_tri3=V3(:,3)*2;
Converting to a single 10-node tetrahedron
dataCell_tri3={nodalData_tri3};
[TRI6,V6,dataCell_tri6]=tri3_tri6(TRI3,V3,dataCell_tri3);
[F6]=element2patch(TRI6,[],'tri6');
nodalData_tri6=dataCell_tri6{1};
Plotting elements
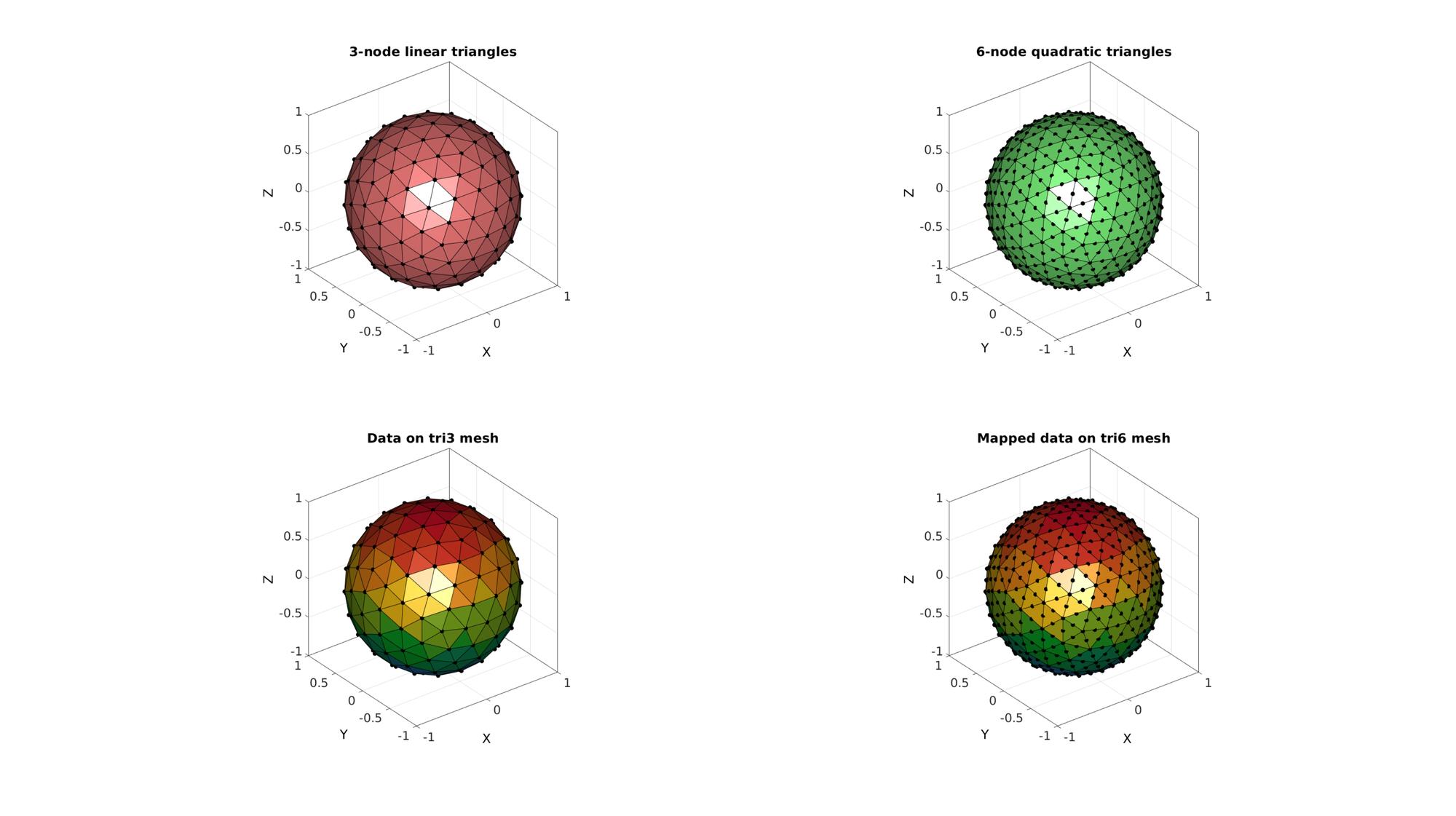
hf=cFigure; % Open figure for plotting subplot(2,2,1); hold on; title('3-node linear triangles','FontSize',fontSize); gpatch(TRI3,V3,'rw'); %Plotting surface plotV(V3,'k.','MarkerSize',25); axisGeom; camlight('headlight'); subplot(2,2,3); hold on; title('Data on tri3 mesh','FontSize',fontSize); gpatch(TRI3,V3,nodalData_tri3); %Plotting surface plotV(V3,'k.','MarkerSize',25); axisGeom; camlight('headlight'); subplot(2,2,2); hold on; title('6-node quadratic triangles','FontSize',fontSize); % gpatch(TRI6(:,[1 4 2 5 3 6]),V6,'gw'); %Plotting surface gpatch(F6,V6,'gw'); %Plotting surface plotV(V6,'k.','MarkerSize',25); axisGeom; camlight('headlight'); colormap(gjet(250)); subplot(2,2,4); hold on; title('Mapped data on tri6 mesh','FontSize',fontSize); gpatch(F6,V6,nodalData_tri6); %Plotting surface plotV(V6,'k.','MarkerSize',25); axisGeom; camlight('headlight'); colormap(gjet(250)); drawnow;

GIBBON www.gibboncode.org
Kevin Mattheus Moerman, [email protected]
GIBBON footer text
License: https://github.com/gibbonCode/GIBBON/blob/master/LICENSE
GIBBON: The Geometry and Image-based Bioengineering add-On. A toolbox for image segmentation, image-based modeling, meshing, and finite element analysis.
Copyright (C) 2019 Kevin Mattheus Moerman
This program is free software: you can redistribute it and/or modify it under the terms of the GNU General Public License as published by the Free Software Foundation, either version 3 of the License, or (at your option) any later version.
This program is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the GNU General Public License for more details.
You should have received a copy of the GNU General Public License along with this program. If not, see http://www.gnu.org/licenses/.